Rosenquist
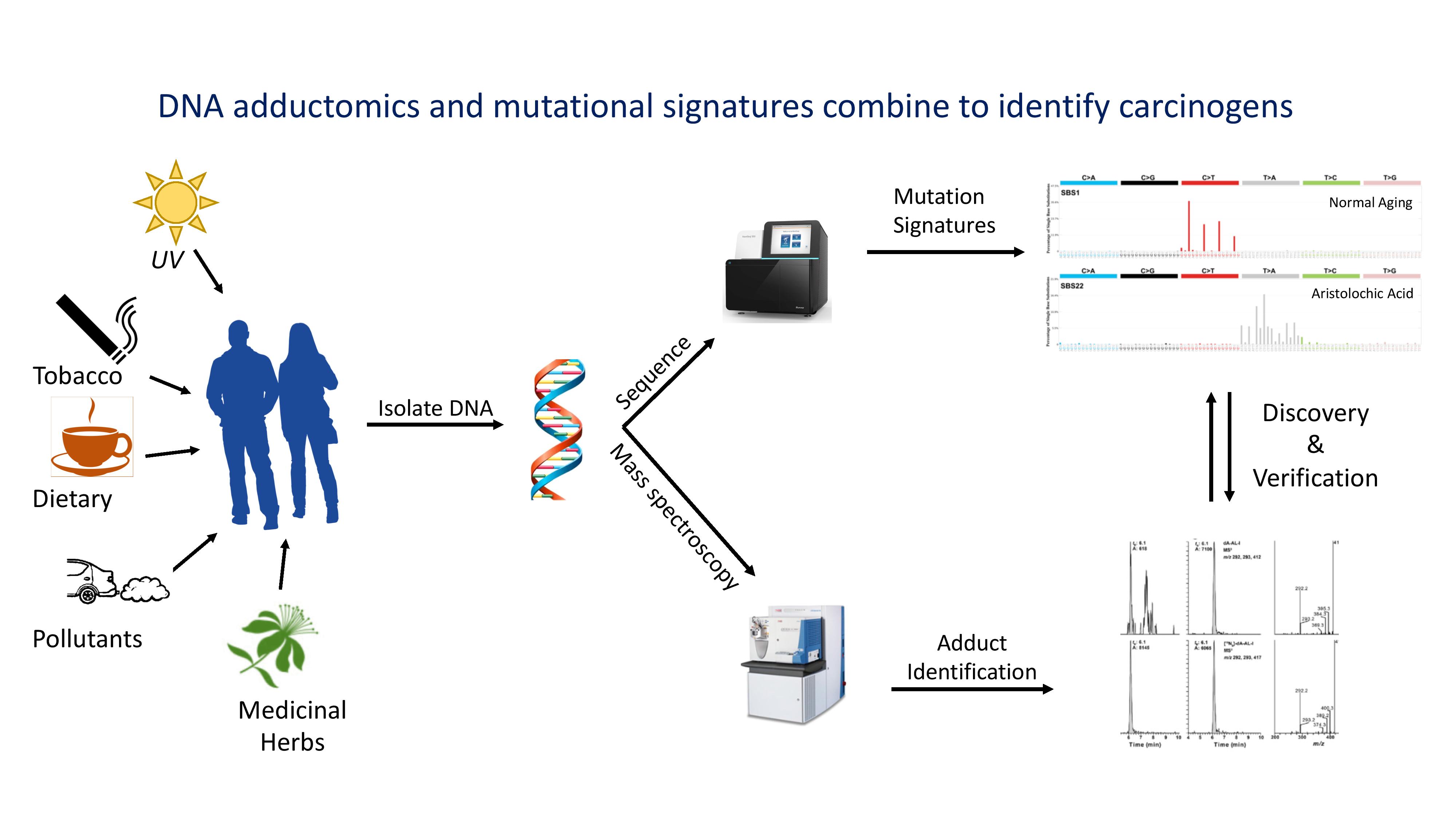
Many cancers are preventable in that they are induced by environmental chemicals. Biomarkers of such exposures are needed to guide public health decisions. We are building on groundbreaking work with the plant carcinogen, aristolochic acid to develop a platform that links damaged DNA bases (adducts) to mutational signatures found in tissue DNA. Specific DNA-adducts, such as those from aristolochic acid and many other chemicals, can be long-lived and serve as excellent biomarkers of exposure. Also, DNA-adducts can leave specific patterns of mutations that are duplicated and maintained in cellular DNA for the lifetime of an individual. Such mutational signatures can provide a diagnostic record of a person’s exposure to mutagenic agents and are evidence of the pathological consequences of those exposures. We are collaborating with University of Minnesota in this approach, called “adductomics” that allow mass-spectroscopic identification and quantification of all the DNA adducts in a patient DNA sample. We have also developed next-gen sequencing analysis to identify the DNA mutations present in single cells of the tissue from which the DNA originated. The goal is to link mutational signatures of unknown etiology to the DNA damage that lead to their generation, and thus to the exposures responsible.
Method for biomonitoring DNA adducts in exfoliated urinary cells by mass spectrometry.
Yun BH, Bellamri M, Rosenquist TA, Turesky RJ.
Anal Chem. Aug 21;90(16):9943-9950 (2018), PMID:30001485
- Supervised mutational sigs predict etiological factors in cancer and reveal the tissue dependence of mutational processes.
Afsari B, Zhang YF, Li L, Kuo A, Danilova L, Favorov A, Rosenquist T, Grollman AP, Kinzler KW, Vogelstein B, Tomasetti C
eLife (2021) 10:e61082 - Targeted and untargeted detection of DNA adducts of aromatic amine carcinogens in human bladder by ultra-performance liquid chromatography-high-resolution mass spectrometry.
Guo J, Villalta PW, Weight CJ, Bonala R, Johnson F, Rosenquist TA, Turesky RJ.
Chem Res Toxicol. Dec 17;31(12):1382-1397 (2018), PMID:30387604 - A rapid throughput method to extract DNA from formalin-fixed paraffin-embedded tissues for biomonitoring carcinogenic DNA adducts.
Yun BH, Xiao S, Yao L, Krishnamachari S, Rosenquist TA, Dickman KG, Grollman AP, Murugan P, Weight CJ, Turesky RJ.
Chem Res Toxicol 30(12):2130-2139 (2017). PMID: 29120619 - Tissues by Ultraperformance Liquid Chromatography-Tandem Mass Spectrometry.
Guo J, Yun BH, Upadhyaya P, Yao L, Krishnamachari S, Rosenquist TA, Grollman AP, Turesky RJ.
Anal Chem. May 3;88(9):4780-7 (2016). PMID:27043225 - Multiclass Carcinogeneic DNA Adduct Quantification in formalin-Fixed Paraffin-Embedded Tissues by Ultraperformance Liquid Chromatography-Tandem Mass Spectrometry.Guo J, Yun BH, Upadhyaya P, Yao L, Krishnamachari S, Rosenquist TA, Grollman AP, Turesky RJ. Anal Chem 88(9):4780-4787(2016). PMID: 27043225
- Mutational signature of aristolochic acid: Clue to the recognition of a global disease.
Rosenquist TA, Grollman AP. DNA Repair 44:205-211(2016). PMID: 27237586 - Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing.
Hoang ML, Kinde I, Tomasetti C, McMahon KW, Rosenquist TA, Grollman AP, Kinzler KW, Vogelstein B, Papadopoulos N.
Proc Natl Acad Sci (2016). PMID: 27528664